Qiang Zhou

Professor Emeritus of Biochemistry, Biophysics and Structural Biology
Lab Homepage: http://mcb.berkeley.edu/labs/zhou/Research Interests
We are studying the mechanisms and factors that control transcriptional elongation and how this control affects HIV replication, latency activation and cancer progression.
Current Projects
The elongation phase of RNA polymerase (Pol) II transcription is widely used to regulate the expression of metazoan genes, especially those for cell growth, renewal and differentiation. Composed of CDK9 and cyclin T (CycT), the positive transcription elongation factor b (P-TEFb) is one of the most important factors that promote the transition of Pol II from promoter-proximal pausing into productive elongation. P-TEFb acts by phosphorylating the Pol II C-terminal domain (CTD) and negative elongation factors to antagonize the latter’s inhibitory effects. This leads to the synthesis of full-length RNA transcripts and the coupling of transcription with pre-mRNA processing.
In human cells, not only
is P-TEFb crucial for the expression of numerous cellular genes, it is also an
indispensable host cofactor for efficient transcription of the HIV genome. Our
recent data indicate that the HIV-encoded Tat protein
and TAR RNA, a stem-loop structure found at the 5’ end of nascent viral
transcripts, recruit P-TEFb and several other transcription
factors/cofactors in the form of the Super Elongation Complex (SEC, Fig. 1) to the integrated HIV proviral genome. SEC contains two different elongation factors, P-TEFb and
ELL2, connected by the scaffold protein AFF4
or AFF1. 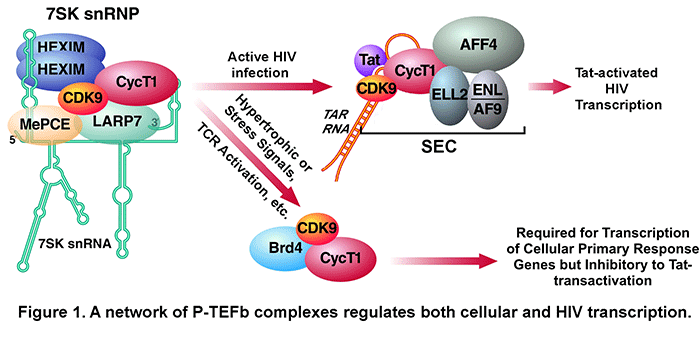 Upon recruitment to the viral promoter, P-TEFb and ELL2 act on
the same polymerase enzyme to synergistically activate HIV transcription. The
sequestration into a SEC and interaction with Tat also stabilize ELL2, which
otherwise would be a short-lived protein targeted by the E3 ubiquitin ligase
Siah1 for proteasomal degradation. Besides being recruited by Tat/TAR to
activate HIV, SEC is also employed by the mixed-lineage leukemia (MLL) protein
and its fusion partners to promote the expression of MLL-target genes and
leukemogenesis.
Upon recruitment to the viral promoter, P-TEFb and ELL2 act on
the same polymerase enzyme to synergistically activate HIV transcription. The
sequestration into a SEC and interaction with Tat also stabilize ELL2, which
otherwise would be a short-lived protein targeted by the E3 ubiquitin ligase
Siah1 for proteasomal degradation. Besides being recruited by Tat/TAR to
activate HIV, SEC is also employed by the mixed-lineage leukemia (MLL) protein
and its fusion partners to promote the expression of MLL-target genes and
leukemogenesis.
In addition to SEC, our data indicate that P-TEFb also exists in at least two other complexes (Fig. 1). First, most cellular P-TEFb is normally sequestered in an inactive state in the 7SK snRNP. We have previously identified 7SK snRNA as well as HEXIM1/2, LARP7 and MePCE proteins as key subunits of this complex and determined their roles in inhibiting CDK9 kinase, maintaining 7SK snRNP integrity and suppressing cellular transformation. A number of conditions/reagents that globally impact cell growth and/or induce stress response can disrupt 7SK snRNP to release P-TEFb for enhanced gene expression. We also show that Tat can directly extract P-TEFb from 7SK snRNP for subsequent assembly of the Tat-SEC complex on HIV promoter.
Another major P-TEFb complex identified by us and others contains the BET bromodomain protein Brd4. This protein acts as the host equivalent of HIV Tat to capture P-TEFb released from 7SK snRNP for delivery to promoters of many cellular, especially primary response genes (Fig. 1) through the interaction of its bromodomains with acetylated histones. Recently, the BET bromodomain inhibitor JQ1 is shown to displace Brd4 from acetylated chromatin thereby inhibiting the transcription of key Brd4/P-TEFb-dependent genes such as c-Myc and inducing differentiation and growth arrest of Myc-dependent cancer cells. As for HIV transcription, because Brd4 is abundant and can directly compete with Tat for binding to P-TEFb, it strongly inhibits Tat-transactivation.
With the identification of three major P-TEFb complexes that constitute a functional network to control HIV and cellular gene expression, we are in a unique position to investigate whether and how this network may be therapeutically exploited to treat human diseases such as HIV/AIDS and cancer. A current project in the lab attempts to manipulate the P-TEFb network to activate transcriptionally silent HIV in latently infected T-cells, which is the primary hurdle to eradication of infection. Identification of agents/strategies that activate latent provirus may, in the presence of highly active antiretroviral therapy (HAART), permit clearance of HIV reservoirs by the immune system. Another project in the lab seeks to identify cancer- and stage-specific changes in the P-TEFb network, which may serve as markers of disease progression and prognosis and potentially be reversed/altered to disrupt oncogenic transformation. We are also performing structure-function analysis of the major P-TEFb complexes to study how they are assembled and how the P-TEFb activity is regulated positively or negatively by its various associated factors.
Selected Publications
The ubiquitin ligase Siah1 controls ELL2 stability and formation of Super Elongation Complexes (SECs) to modulate gene transcription. [M. Liu, J. Hsu, C. Chan, Z. Li, and Q. Zhou (2012) Mol Cell, 46, 325-334]
RNA Polymerase II elongation control. [Q. Zhou, T. Li, and D.H. Price (2012) Annu. Rev. Biochem. 81, 119-143]
Human Polymerase-Associated Factor complex (PAFc) connects the Super Elongation Complex (SEC) to RNA polymerase II on chromatin. [N. He, C.K. Chan, B. Sobhian, Y. Xue, M. Liu, M. Benkirane, and Q. Zhou (2011) Proc. Natl. Acad. Sci. USA.108, 14719-14720]
The control of HIV transcription: Keeping RNA Polymerase II on track. [M. Ott, M. Geyer, and Q. Zhou (2011) Cell Host & Microbe 10, 426-435]
HIV-1 Tat and host AFF4 recruit two transcription elongation factors into a bifunctional complex for coordinated activation of HIV-1 transcription. [N. He, M. Liu, J. Hsu, Y. Xue, S. Chou, A. Burlingame, N.J. Krogan, T. Alber, and Q. Zhou (2010). Mol Cell 38, 428-438]
A La-related protein modulates 7SK snRNP integrity to suppress P-TEFb-dependent transcriptional elongation and tumorigenesis. [N. He, N.S. Jahchan, E. Hong, M. A. Bayfield, R. J. Maraia, K. Luo, and Q. Zhou (2008) Mol. Cell 29, 588-599]
Brd4 recruits P-TEFb to chromosomes at late mitosis to promote G1 gene expression and cell cycle progression. [Z. Yang, N. He and Q. Zhou (2008) Mol. Cell. Bio. 28, 967-976]
PP2B and PP1α cooperatively disrupt 7SK snRNP to release P-TEFb for transcription in response to Ca2+ signaling. [C. Chen, M. Liu, H. Li, Y. Xue, W.N. Ramey, N. He, N. Ai, H. Luo, Y. Zhu, N. Zhou, and Q. Zhou (2008) Genes & Dev.22, 1356-1368]Tat competes with HEXIM1 to increase the active pool of P-TEFb for HIV-1 transcription. [M. Barboric, J. H. N. Yik, N. Czudnochowski, Z. Yang, R. Chen, X. Contreras, M. Geyer, B. M. Peterlin, and Q. Zhou (2007) Nuc. Acids Res. 35, 2003-2012]
Recruitment of P-TEFb for stimulation of transcriptional elongation by bromodomain protein Brd4. [Z. Yang, J. H. N. Yik, R. Chen, M. K. Jang, K. Ozato and Q. Zhou (2005) Mol. Cell 19, 535-545]
Inhibition of P-TEFb (CDK9/cyclin T) kinase and RNA polymerase II transcription by the coordinated actions of HEXIM1 and 7SK snRNA. [J. H. N. Yik, R. Chen, R. Nishimura, J. L. Jennings, A. J. Link and Q. Zhou (2003) Mol. Cell 12, 971-982]
The 7SK small nuclear RNA inhibits the Cdk9/cyclin T1 kinase to control transcription. [Z. Yang, Q. Zhu, K. Luo, and Q. Zhou (2001) Nature 414, 317-322]
Stimulatory effect of splicing factors on transcriptional elongation. [Y. Fong and Q. Zhou (2001) Nature 414, 929-933]
Photo credit: Mark Hanson of markjosephstudio.com.
Last Updated 2012-07-05