Jennifer A. Doudna

Howard Hughes Medical Institute Investigator, Li Ka Shing Chancellor's Chair in Biomedical and Health Sciences, Professor of Molecular Therapeutics
Lab Homepage: https://doudnalab.orgResearch Interests
RNA molecules are uniquely capable of encoding and controlling the expression of genetic information, often as a consequence of their three-dimensional structures. We are interested in understanding and harnessing RNA-mediated control of the genome, including CRISPR-Cas bacterial adaptive immunity and related systems.
Current Projects
 The CRISPR bacterial adaptive immune system
The CRISPR bacterial adaptive immune system
Prokaryotes have evolved a nucleic acid-based immune system that shares some functional similarities with RNA interference in eukaryotes. Central to this system are DNA repeats called CRISPRs (Clustered Regularly Interspaced Short Palindromic Repeats). CRISPRs are genetic elements containing direct repeats separated by unique spacers, many of which are identical to sequences found in phage and other foreign genetic elements. Recent work has demonstrated the role of CRISPRs in adaptive immunity and shown that small RNAs derived from CRISPRs (crRNAs) are implemented as homing oligos for the targeted interference of foreign DNA.
Phylogenetic analysis of CRISPR-associated (Cas) proteins suggests there are at least seven distinct versions of this immune system. These systems can be extremely divergent mechanistically and provide a rich area to research RNA:protein interactions, including novel protein folds. To explore this diversity, we utilize metagenomic analysis of microbial communities across the tree of life to discover novel proteins for efficient genome editing. These systems are tested for function in vitro, heterologously in vivo, in situ microbial communities, and in eukaryotic cells.
DNA Acquisition in the CRISPR-Cas system RNA-guided RNA cleavage by the Csm complex
CRISPR-harboring organisms generate immunological memory of previous infections by capturing short segments of foreign DNA for integration into CRISPR loci as spacer sequences. Central to this process are Cas1 and Cas2 – which are generally conserved in most CRISPR systems. Using a combination of biochemical, structural, and genetic approaches, we found that Cas1 and Cas2 functions as a protein complex. The E.coli Cas1–Cas2 complex (Type IE) captures ~30 bp of foreign DNA and integrates them into the CRISPR locus via a direct nucleophilic reaction similar to many retroviral integrases and transposases. Our results uncover the structural basis for foreign DNA capture and the mechanism by which Cas1–Cas2 functions as a molecular ruler to dictate the sequence architecture of CRISPR loci. We have also demonstrated how an integration host factor helps Cas1-Cas2 recognize the CRISPR array for site-specific integration. Recently, we found that a mini-integrase comprising the type V-C Cas1 protein alone catalyzes DNA integration with a shortened ruler mechanism, which may give insight to the function of the ancestral Cas1, prior to Cas2 adoption.
Engineering CRISPR-Cas Proteins
The Doudna lab employs protein engineering and screening strategies to bestow CRISPR-Cas enzymes with improved and novel functionalities. In collaboration with Stanley Qi and the laboratories of Wendell Lim, Jonathan Weissman, and Adam Arkin, we pioneered the engineering of CRISPR-Cas proteins for new purposes by converting Cas9 into a tool for targeted control of gene expression, referred to as CRISPR interference (CRISPRi). In this platform, catalytically inactive Cas9 serves as an RNA-guided DNA binding protein to silence gene expression in a sequence-specific manner. In collaboration with the laboratory of David Savage, we have also developed a series of high-throughput mutant library construction, screening, and sequencing pipelines to probe CRISPR-Cas protein function and to isolate engineered variants with novel properties. In one example, we utilized domain insertion profiling with sequencing (DIP-seq) to identify an allosteric Cas9 switch, yielding an optimized Cas9 variant that is activated by the small molecule 4-hydroxytamoxifen. Recently, we used a similar large-scale protein engineering and screening platform to construct circularly permuted, viral protease-activated ProCas9s that orchestrate programmed cellular responses to pathogen-associated protease activity.
Nucleic Acid Targeting CRISPR-Cas systems
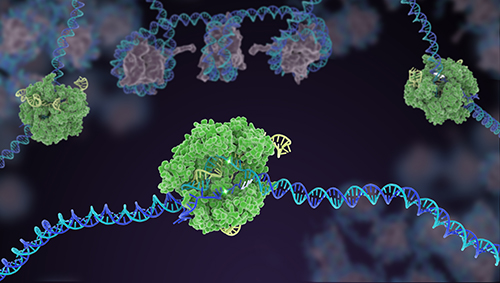
We employ traditional biochemistry, combined with sequencing, live-cell imaging, and structural biology approaches, to uncover the fundamental mechanisms driving the CRISPR effectors now being developed as genome editing and diagnostic tools.
RNA-targeting CRISPR-Cas systems - We use biochemical activity assays in conjunction with structural techniques and sequencing to investigate RNA-targeting CRISPR systems, such as the Type VI effector, Cas13a. A crystal structure of L. bacterium Cas13a bound to a single guide RNA (sgRNA) demonstrated, together with biochemical studies, how crRNAs are processed by this enzyme and how catalytic activity is blocked before target binding. (Knott et al., 2017) We also use high-throughput sequencing, together with biochemistry, to discover mechanisms governing RNA-targeting CRISPR effectors, such as the decoupled RNA targeting and HEPN nuclease activity in LbaCas13a. (Tambe et al, 2018)
DNA-targeting CRISPR-Cas systems - Biochemistry allows us to investigate the mechanisms driving DNA-targeting CRISPR effectors on a molecular level. These studies uncovered indiscriminate, single-stranded DNAase activity in Cas12a unleashed by target binding, the chemistry behind the DETECTR technology (vide infra). (Chen et al., 2018) Structural (Cryo-EM), biochemical, and live-cell studies, as part of a collaboration with the laboratory of Eva Nogales, moreover characterized the fundamental properties of distinct, DNA-targeting CRISPR-CasX effectors. (Liu, Orlova, Oakes, et al., 2019) These studies, in combination with techniques such as high-throughput sequencing and live-cell imaging, provide important insight that helps better engineer and control CRISPR genome editing tools.
Dual RNA- and DNA-targeting CRISPR-Cas systems - We use a combination of biochemical and structural biology techniques (single-particle EM) to investigate the mechanism and function of dual DNA- and RNA-targeting Type III CRISPR-Cas systems. In collaboration with the laboratories of Eva Nogales, Akeo Shinkai, and John van der Oost, we have revealed the molecular architecture and RNA targeting mechanism of T. thermophilus Type III-B CRISPR complexes. In collaboration with Eva Nogales’ lab, we are also investigating the pathway of transcription-coupled DNA and RNA targeting by Type III-A CRISPR-Cas systems. Insights into how Type III interference complexes recognize and degrade their targets may lead to improved tools for controlling gene expression in vivo.
Editing in disease: CRISPR-Cas gene editing for the study and treatment of human diseases
CRISPR-Cas is a gene-editing technology that has the potential to dramatically reshape the treatment of genetic diseases by replacing or deleting problematic genes. Cas proteins can be delivered non-genetically in the form of a ribonucleoprotein (RNP) complex - one method for direct and efficient gene editing in cells - our laboratory has recently developed Cas9 RNPs that are cell penetrant in the context of the adult mouse brain. We are currently exploring therapeutic use of Cas9 as a method to treat Huntington’s Disease (HD). HD is a neurodegenerative disorder caused by a genetically dominant, CAG trinucleotide expansion in the Huntingtin (HTT) gene, and suppression of mutant Huntington protein (mHTT) is currently being explored as a therapeutic method to treat HD. We are using Cas9 RNP technology to reduce the levels of mHTT, with the goal of halting the primary disease mechanism and progression of HD. Along with Huntington’s disease, we are also interested in using CRISPR-based technologies to study and treat various other brain diseases. In particular, we are focusing our efforts on primary glioblastoma (GBM), which is both the most common and most devastating form of malignant brain cancer in adults. Current treatment options are limited, with GBM progression leading to death within 2 years of diagnosis in 90% of cases. In collaboration with the Costello lab at UCSF and the Fellmann lab at the Gladstone Institutes, we are using a variety of CRISPR strategies including Cas9 circular permutants and cell-penetrating Cas9 ribonucleoproteins to understand and exploit genetic vulnerabilities of GBM.
Delivery of CRISPR-Cas tools to disease-relevant human cells
The therapeutic use of CRISPR-Cas requires the effective transportation of gene editing tools into disease-relevant cells and tissues. We therefore are developing strategies for the effective delivery of gene editing tools for both ex vivo and in vivo use in mammals. Recently, our laboratory has generated cell-penetrating Cas9 ribonucleoproteins by the iterative addition of nuclear-localization signals. These cell-penetrating Cas9 ribonucleoproteins are effective at mediating gene editing in the neuronal cells of mice and are a new strategy for directly altering disease-causing genes in the brain. We collaborate closely with the UC Berkeley laboratories of Matt Francis and Ross Wilson to design and test both viral and non-viral strategies for the delivery of CRISPR-Cas tools. We are building upon our initial successes to design new strategies for the effective delivery of Cas9 ribonucleoproteins.
Selected Publications
CasX enzymes comprise a distinct family of RNA-guided genome editors. Liu JJ, Orlova N, Oakes BL, Ma E, Spinner HB, Baney KLM, Chuck J, Tan D, Knott GJ, Harrington LB, Al-Shayeb B, Wagner A, Brötzmann J, Staahl BT, Taylor KL, Desmarais J, Nogales E, Doudna JA. (2019) Nature.
Programmed DNA destruction by miniature CRISPR-Cas14 enzymes. Harrington LB, Burstein D, Chen JS, Paez-Espino D, Ma E, Witte IP, Cofsky JC, Kyrpides NC, Banfield JF, Doudna JA. (2018) Science.
CRISPR-Cas12a target binding unleashes indiscriminate single-stranded DNase activity. Chen JS, Ma E, Harrington LB, Da Costa M, Tian X, Palefsky JM, Doudna JA. (2018) Science.
Enhanced proofreading governs CRISPR-Cas9 targeting accuracy. Chen JS, Dagdas YS, Kleinstiver BP, Welch MM, Sousa AA, Harrington LB, Sternberg SH, Joung JK, Yildiz A, Doudna JA. (2017) Nature.
Integrase-mediated spacer acquisition during CRISPR-Cas adaptive immunity. Nuñez JK, Lee AS, Engelman A, Doudna JA. (2015) Nature.
Enhanced homology-directed human genome engineering by controlled timing of CRISPR/Cas9 delivery. Lin S, Staahl BT, Alla RK, Doudna JA. (2014) Elife.
Programmable RNA recognition and cleavage by CRISPR/Cas9. O'Connell MR, Oakes BL, Sternberg SH, East-Seletsky A, Kaplan M, Doudna JA. (20140) Nature.
DNA interrogation by the CRISPR RNA-guided endonuclease Cas9. Sternberg SH, Redding S, Jinek M, Greene EC, Doudna JA. (2014) Nature.
RNA-programmed genome editing in human cells. Jinek M, East A, Cheng A, Lin S, Ma E, Doudna JA. (2013) Elife.
A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Jinek M, Chylinski K, Fonfara I, Hauer M, Doudna JA, Charpentier E (2012) Science.
Photo credit: Keegan Houser.
Last Updated 2021-06-17