By Shana Singh 2016
For decades, three-dimensional structures of proteins and protein complexes have been determined by X-ray crystallography or NMR spectroscopy. In 1952, X-ray crystallography was used by Rosalind Franklin to generate the crucial first image of DNA, enabling Watson and Crick to use it as the basis for their double helix model. More recently, NMR spectroscopy has been used by researchers to create computer models of small proteins. Even though many advances have been made, these techniques have their limitations, as many proteins are nearly impossible to crystallize (inhibiting the use of X-ray crystallography) or may be too large to be solved with NMR.
Early Work with X-Ray Crystallography...
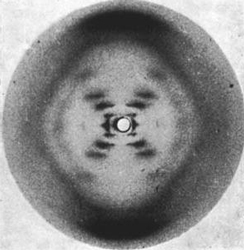
Rosalind Franklin (courtesy of Jewish Chronicle Archive/Heritage-Images) and an image from her x-ray crystallography research (courtesy of Raymond Gosling/King's London).
Today, technical advances in single particle cryo-electron microscopy (cryo-EM) have drastically increased our ability to understand protein structures by addressing some of the limitations of X-ray crystallography and NMR Spectroscopy. Because of its enormous potential to revolutionize science, cryo-EM was named “Method of the Year” in 2015. Researchers at Berkeley currently have access to two cryo-EM instruments, one at LBNL (JEOL 3100) and one at QB3 (Titan), both with direct detectors. However, new advances can allow for facilities with much higher throughput -- enabling research on a much larger scale while still maintaining a high degree of quality.
With innovation comes great responsibility, and because this new technology has such a high price tag, there was concern that only researchers within private and highly-funded facilities would have access to it. To help level the playing field, MCB Professor Eva Nogales and a group of researchers from four UC campuses and the Lawrence Berkeley National Laboratory have now secured $10 million in HHMI funds to create a regional facility for this pioneering cryo-EM technology in the Bay Area. This new facility will provide access to this cutting-edge technology to the participating institutions in the Bay Area (UC Berkeley, UC Davis, UC Santa Cruz, UCSF and LBNL).
Using Today's Technologies...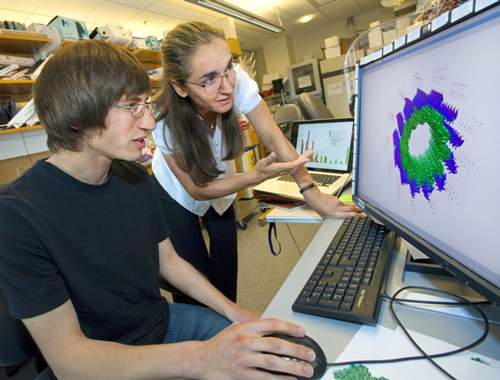
Gregory Alushin and Eva Nogales studying images of microtubule structures. (Photo by Roy Kaltschmidt)
3D Cryo-EM can solve the mysteries of enigmatic protein structures, which will ultimately lead to a mechanistic understanding of how they work in the cell. Using high-end electron microscopes and new direct detectors, enough data can be obtained to construct density maps. Researchers can then use these maps to generate atomic models of protein structures. In other words, the new cryo-EM instrumentation provides the clarity of protein structure that was elusive for decades before! Cryo-EM will have a profound impact on the way researchers study many different aspects of molecular biology, including genome editing, translation, autophagy, pluripotency, chromatin structure, and more.
Cryo-EM structure of the microtubule showing atomic details of the contact between tubulins, the protein building blocks.
The Bay Area will soon have not one but two “multi-user, high throughput cryo-EM” facilities (one at the UC Berkeley campus and one in Mission Bay). These two sites will provide greater opportunities for collaboration than ever before, ensuring that the Bay Area remains a hub of exploration and discovery in the field of structural biology.