Xavier Darzacq

Edward E. Penhoet Distinguished Endowed Chair in Global Health and Infectious Disease, Professor of Molecular Therapeutics*
*and
Research Interests
Transcription regulation during cellular differentiation. Linking the biophysical rules of nuclear organization and gene expression control mechanism.
Current Projects
Our group studies transcription focusing on the role imposed by nuclear architecture on the molecules regulating it. We use a model system of cellular differentiation of human primary dermal fibroblasts into myofibroblasts, which is relevant to wound healing. Among the specificities of this system, we show that the maintenance of the fibroblast state depends on a specialized non-canonical mediator complex, which is becoming one of the central projects of the group. Moreover several transcription factors equally expressed in the two states are required for the differentiation process and we will focus our research on the regulation of their activity as transcriptional regulators.
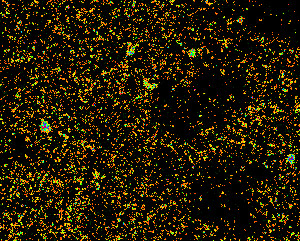
Clusters or RNA polymerase II in a human cell nucleus
Over the last five years we developed new techniques to study the organization of proteins in the nucleoplasm with resolutions in the range of a few nanometers as well as procedures to track single molecules in live cells in order to measure their specific and unspecific interactions with chromatin as well as their space exploration properties. Together with more established techniques such as fluorescent in situ hybridization (FISH), live cell visualization of nucleic acids (Lac, Tet and MS2) or photobleaching techniques (FRAP, FLIP), we can image transcription regulation with the appropriate temporal resolution (a few milliseconds to a few days) and at single molecule resolution within live cells.
Our recent findings suggest a very strong link in between the organization of the nucleoplasm and the biophysical rules governing macromolecular assemblies. While a decade ago the fast mobility of proteins in the nucleoplasm lead to diffusion models, we are now envisioning models where proteins are guided on DNA or protein networks controlling their exploration properties (diffusion can be guided). We are currently deeply involved in providing evidences for such a model by characterizing the nucleoplasmic distribution of chromatin and non-chromatin components of the nucleus as well as measurements of single molecules explorations and binding.
Within the next decade, great technological advances are expected in the field of microscopy, fluorophore chemistry as well as cellular engineering with new genome editing techniques already in place. These new developments will put us in an ideal situation to monitor several different molecules simultaneously (therefore defining molecular multicomponent complexes) with resolutions close to that of electron microscopy. It will be a unique opportunity to integrate the cellular and biochemical know how of an institute strong of a long experience in gene regulation with advances in biophysics, chemistry and imaging in order to reveal the rules governing transcription regulation within a living cell and eventually living organisms.
Selected Publications
Senecal A, Munsky B, Proux F, Ly N, Braye FE, Zimmer C, Mueller F and Darzacq X. (2014) Transcription Factors Modulate c-Fos Transcriptional Bursts. Cell Reports, (in press DOI: 10.1016)
Woringer M, Darzacq X, Izeddin I. (2014) Geometry of the nucleus: a perspective on gene expression regulation. Current Oppinion in Chemical Biology, 20, 112-19
Izeddin I, Récamier V, Bosanac L, Cisse II, Boudarene L, Dugast-Darzacq C, Proux F, Bénichou O, Voituriez R, Bensaude O, Dahan M, Darzacq X. (2014) Single-molecule tracking in live cells reveals distinct target-search strategies of transcription factors in the nucleus. Elife, 10.7554/eLife.02230.
Récamier V., Izeddin I., Bosanac L., Dahan M., Proux F. and Darzacq X. (2014) Single cell correlation fractal dimension of chromatin: a framework to interpret 3D single molecule super-resolution Nucleus, 5, 75-84
Cisse I.I., Izzedin I., Causse S.Z., Boudarene L., Muresan L., Dugast-Darzacq C., Hajj B., Dahan M., Darzacq X. (2013) Real time dynamics of RNA Polymerase II clustering in live human cells. Science 341, 664-7
News & Views : Rickman C and Bickmore WA, Flashing a light on the spatial organization of transcription. Science, 341, 621-2.
Haimovich G., Medina D., Causse S.Z., Garber M., Millán-Zambrano G., Barkai O., Chávez S., Pérez-Ortín J.E., Darzacq X. and Choder M. (2013) Gene expression is circular: factors for mRNA degradation also foster mRNA synthesis Cell, 153, 1000-11
Mueller F., Sénécal S., Tantale K., Marie-Nelly H., Ly N., Collin O., Basyuk E., Bertrand E., Darzacq X., Zimmer C. (2013) FISH-QUANT: automated counting of single mature and nascent transcripts in 3D FISH images. Nature Methods. 10, 277-278 34
Normanno D., Dahan M., Darzacq X. (2012) Intra-nuclear mobility and target search mechanisms of transcription factors: A single-molecule perspective on gene expression. BBA-Gene Regulatory Mechanisms. 1819(6):482-93
Izeddin, I., El Beheiry, M., Andilla, J., Ciepielewski, D., Darzacq, X., Dahan, M. (2012) PSF shaping using adaptive optics for three-dimensional single-molecule super-resolution imaging and tracking. Optics Express 20: 4957-4967
Darzacq X., Yao J., Larson D., Causse S.Z., Bosanac L., deTurris V., Ruda V.M., Lionnet T., Zenklusen D., Guglielmi B., Tjian R. and Singer R.H. (2009) Imaging Transcription in Living Cells. Annual Review of Biophysics 38, 173-196
Darzacq X., Shav-Tal Y, de Turis, V. Brody, Y., Shenoy, S.M., Phair, R.D., Singer R.H. (2007) In Vivo Dynamics of Polymerase II Transcription. Nat Struct Mol Biol 14, 796-806 (Selected Article of the month by NSMB)
News & Views : Lamond A.I., Swedlow J.R. (2007) RNA polymerase II transcription in living color. Nat Struct Mol Biol. 14, 788-790.
Shav-Tal Y, Darzacq X., Shenoy, S.M., Fusco D., Janicki S.M., Spector D.L., Singer R.H. (2004) Dynamics of Single mRNPs in Nuclei of Living Cells. Science, 304,1797-80
Photo credit: Mark Hanson at Mark Joseph Studios.
Last Updated 2014-07-02