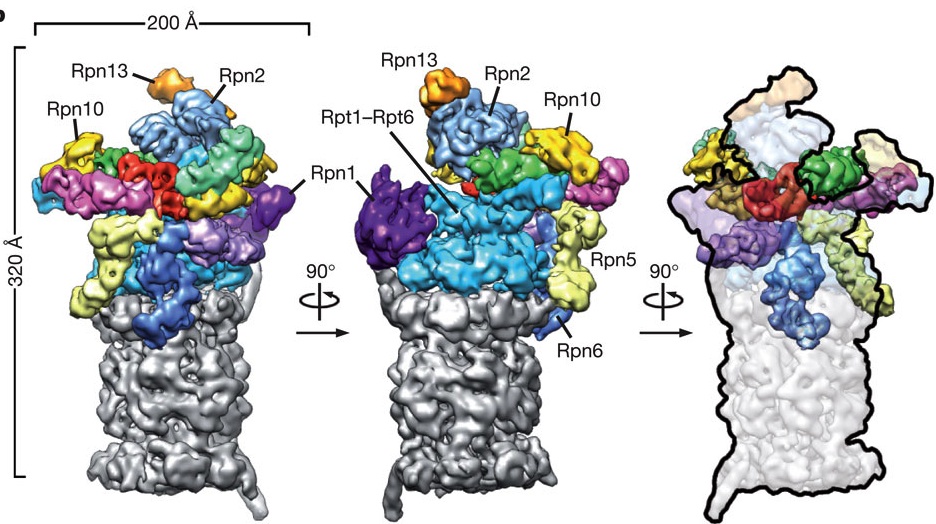
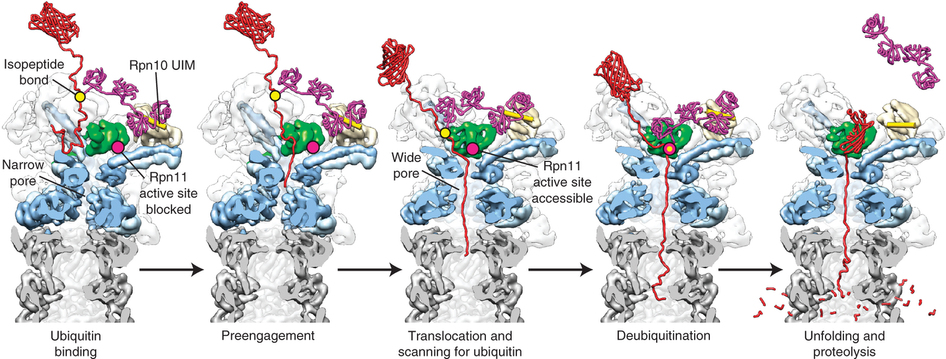
Studying the 26S proteasome
Our goals are to decipher the fundamental principles that govern substrate engagement, de-ubiquitylation, unfolding, and translocation by the eukaryotic 26S proteasome.
In all cells, the degradation of intracellular proteins is highly specific and tightly regulated by ATP-dependent compartmental proteases. These proteases belong to the AAA+ ATPase family, whose characteristic feature is a structurally conserved ATPase domain that assembles into oligomeric rings and converts the energy of ATP binding/hydrolysis into conformational changes to perform mechanical work. Molecular machines of this enzyme family are also active, for instance, as helicases, translocases, clamp loaders, or motor proteins. For protein degradation, the AAA+ ATPases collaborate with compartmental peptidases. In these complexes, the ATPases recognize appropriate protein substrates and harness ATP hydrolysis to drive the mechanical unfolding and the translocation of the polypeptide chain into a sequestered degradation chamber of the associated peptidase.
The research of my lab focuses on the 26S proteasome, the major ATP-dependent protease in eukaryotic cells. In addition to processing misfolded, damaged, and incompletely translated proteins, the proteasome selectively degrades a large variety of regulatory proteins that are involved in cell-cycle control, cell differentiation, signal transduction, transcription, and other vital processes, making it a very important mediator of post-translational control.
Most proteasomal substrates are marked for degradation by the attachment of a poly-ubiquitin chain, which acts as a tethering signal for substrate delivery. Although substantial knowledge about the ubiquitin-tagging systems has already been accumulated, the mechanisms underlying proteasomal substrate degradation, the irreversible and hence critical step for the unidirectional control of cellular pathways, remain largely elusive.