Curiosity Driven Basic Research, Education, Training and a Commitment to Public Service
Welcome
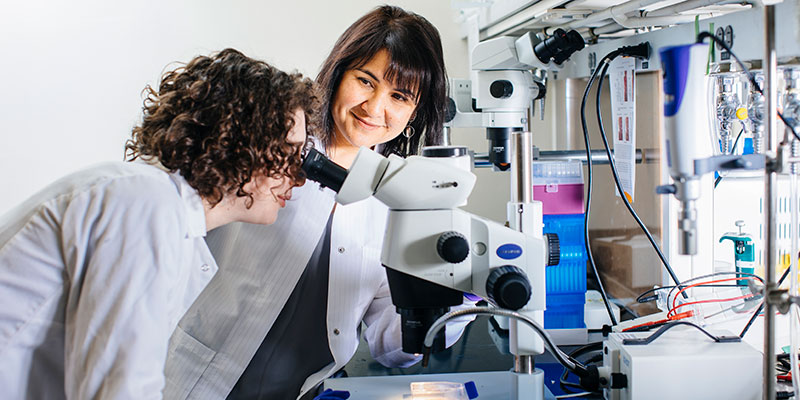
The teaching and research activities of the Department of Molecular and Cell Biology (MCB) concern the molecular structures and processes of cellular life and their roles in the function, reproduction, and development of living organisms. The types of living organisms from which the departmental faculty draws its working materials range from viruses and microbes through plants, roundworms, annelids, arthropods, and mollusks to fish, amphibia, and mammals.



 Biochemistry, Biophysics
Biochemistry, Biophysics Immunology and
Immunology and Cell Biology, Development
Cell Biology, Development
 Genetics, Genomics,
Genetics, Genomics,



